This time of year I have to file my renewal for my DOE Fellowship, including a statement of accomplishments and a summary of the computational aspects of my work. Seemed like a good idea to jot these down in my notebook as well:
synopsis of accomplishments over the year:
In January I passed my qualifying exam, advancing to candidacy where my dissertation remains my only requirement. In March I attended the phylogenetics workshop in Bodega Bay, an internationally known week-long training program. I also attended the CSGF conference in June, and gave talks at both the Evolution conference and the integrative phylogenetics conference, iEvoBio, in July, both key events in my field.
From the beginning of June until the start of September, I stepped outside my research in phylogenetic models to focus on stochastic population dynamics. While the details of this project are described in my practicum evaluation, the mathematical and computational tools I learned in the process have found surprisingly apt applications in my phylogenetics research, where they have helped me solve a key problem in my thesis. Since this breakthrough, I have been focused on writing up two manuscripts which have been made possible by this step forward.
As my computational needs surpass what is readily available through my lab servers, I have learned to leverage Amazon’s cloud computing for rapid deployment of large-scale simulations and analyses. Overall, this has been a productive year, translating what I’ve learned in my POS classes into practical research experience and making some important steps in my thesis. I look forward to the next year of synthesizing and writing up these results and continuing to build on the methods and tools I’ve learned and developed.
describe concisely what you regard as the most interesting and innovative aspects of your current research, with special emphasis on computational science.
My research combines the recent explosion of data available through gene sequencing and subsequent readily available phylogenetic information with the power of high-performance computational approaches to “replay the tape” of evolution to better understand the origins and maintenance of biodiversity.
Genetic data paved the way to phylogenetic inference, the mapping of ancestor-descendant relationships between the species living today. These phylogenies become my periscope into the past. In the absence of fossils, I can simulate different evolutionary models of selection on various traits retracing the phylogenetic tree to discover which models are most consistent with the patterns in species we observe today. By running the stochastic simulation repeatedly, I can create not only one possible outcome, but the parameter space of all possible outcomes and their associated probabilities. This planet has run the experiment of evolution only once, but using this computationally intensive approach I can run that process over and over again under different assumptions, allowing me to explore evolutionary questions that can be answered no other way.
My own reflections and list
Hmm, the research summary with computational emphasis is kinda vague, but aiming for a non-specialist audience. Hard to differentiate between the aspects that really leverage HPC and those that can be adequately run at the desktop level. Is that really the best statement of my accomplishments? I might try a different format for myself:
Formulated and explored implications of my phylogenetic Monte Carlo approach. Formulated throughout the Spring quarter, been writing and rewriting the manuscript most of this past Fall.
Derived and implemented the heterogeneous alpha/sigma model (in a few busy days and a long train ride in July, waiting to write up after PMC is done)
Used said model and phylogenetic Monte Carlo to demonstrate clear release-of-constraint in the Labrid data
Exploring better early warning signal models and detection methods. Settled on linearized models of the canonical form of two bifurcation types, identifying warning signals directly by a similar Monte Carlo test to the phylogenetics example.
Explored stochastic population dynamics in structured populations via the ELPA tribolium model. Negative feedbacks mean no large-amplification demographic noise, but cycles do break down the linear noise approximation. (a few weeks in May and June)
Rebuilt the Adaptive Dynamics code to illustrate how time to branching and coexistence times vary in parameter space (auto-generation of the butterfly plots, mostly two weeks in April)
Began my open lab notebook (starting in January on OWW, migrated to Wordpress in Sept), public code hosting (initially google code, moved to github in March), Mendeley database (Oct ’09), Flickr only this Oct, and social media for science (Twitter only since July, though have used FF since the beginning of the year).
Started AWS/cloud computing. Still coding in C but using R more, parallelization in R’s snowfall.
In this spirit, Mendeley provides some easy data on what I’ve been reading:
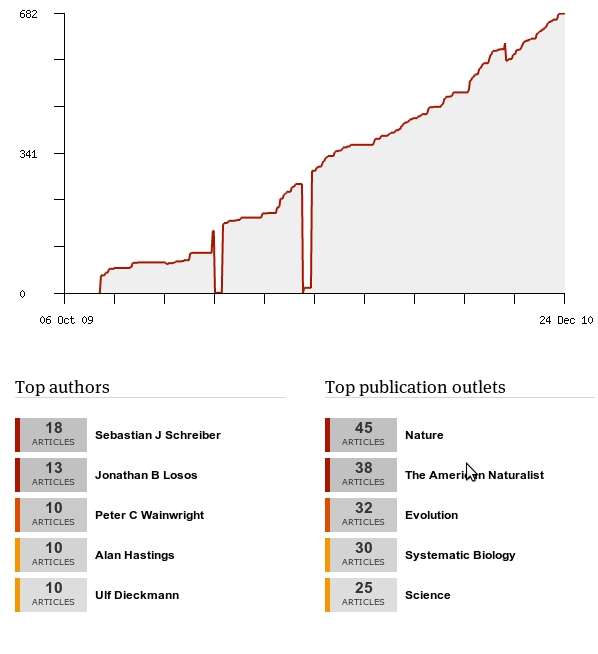
Interesting if not too surprising that my most popular authors are all advisors of mine, except Losos. (Also it’s really obvious where I twice deleted my Mendeley database and needed their help support to recover my data… in only a few days).
.