Summarizing parrotfish data under the single shift at the intramandibular joint. See version stable code for data loading/transforms (parrotfish_data.R), and for plots (parrotfish.R) as shown, and for analysis functions (loop_models_traits_regimes.R). Models are named by the parameter which is regime-dependent. Results are shown sorted by likelihood, so the best model is on top, and likelihood values are scaled by the weakest model.
Tree: 
alpha is favored in all the ratio traits, sometimes substantially. 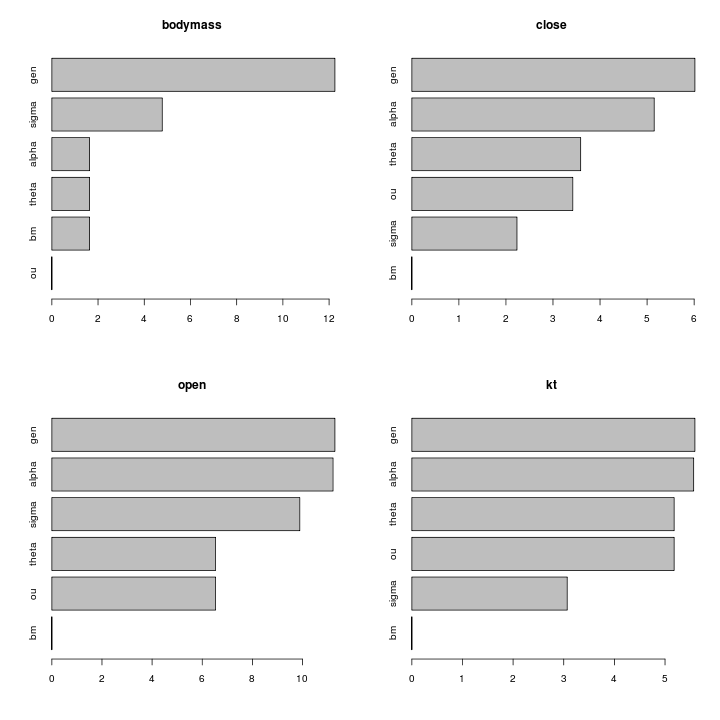
Without size corrections the muscle traits essentially reflect body mass. Though ratio may be better than the regression:
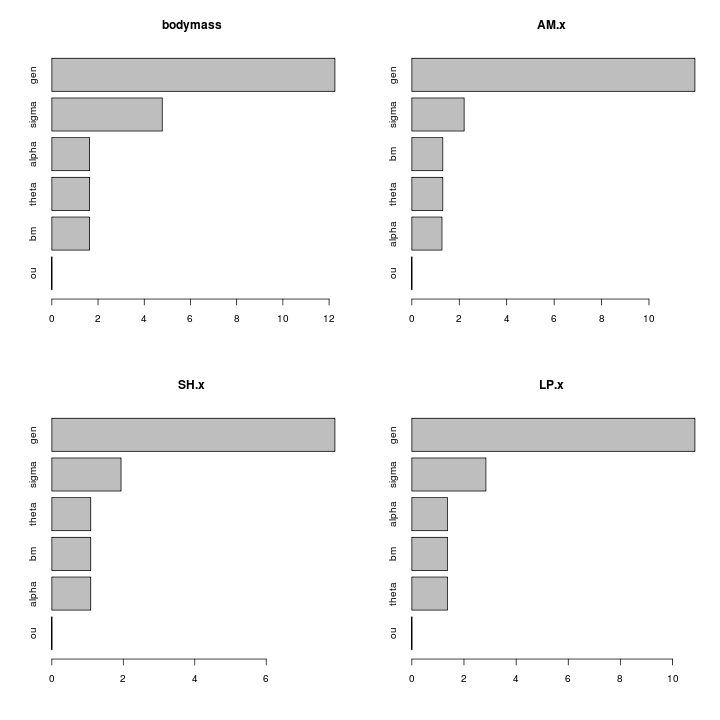
Size-corrected length traits show a substantial signal:
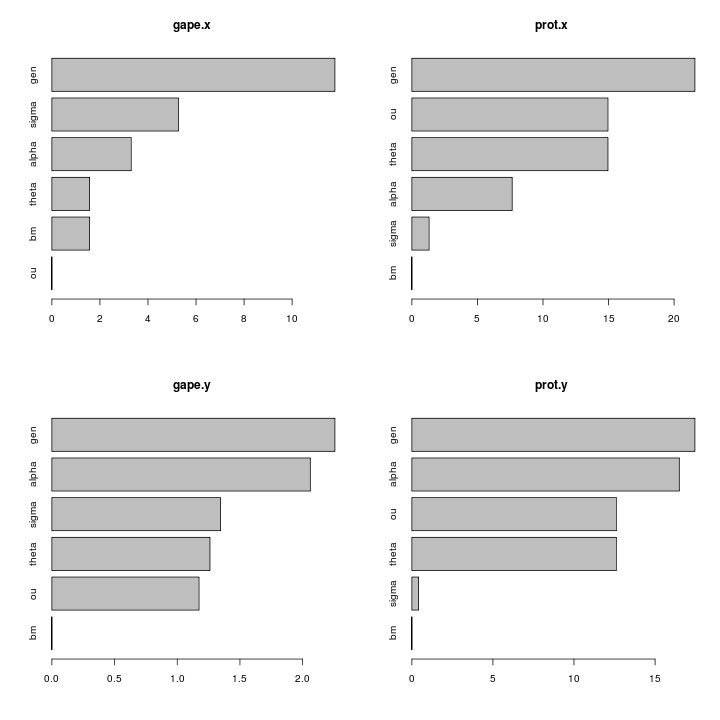
The appaxial muscle looks strong candidate for release of constraint, the control muscles not involved with the jaw mechanism do not show any such release:
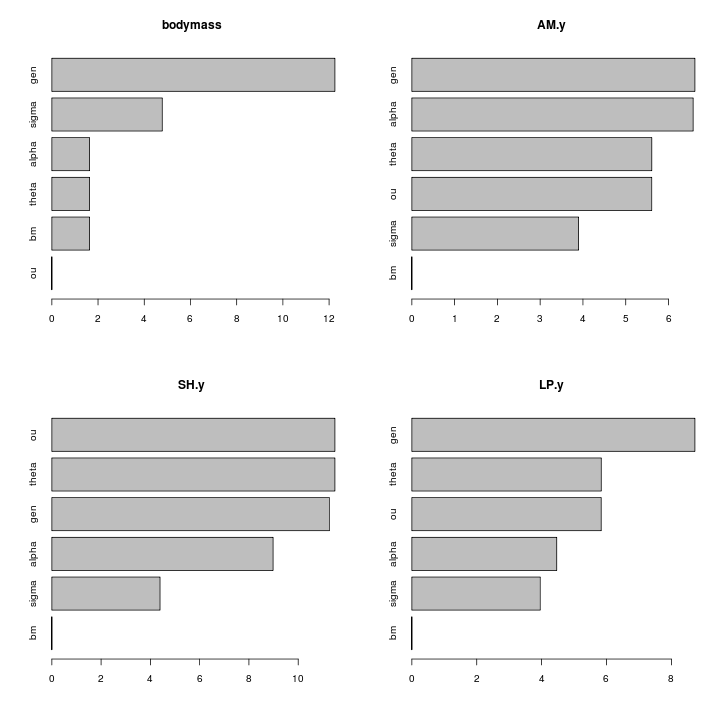
Strength of constraint (alpha parameter), by trait: log ratio of intramandibular to other parrotfish: 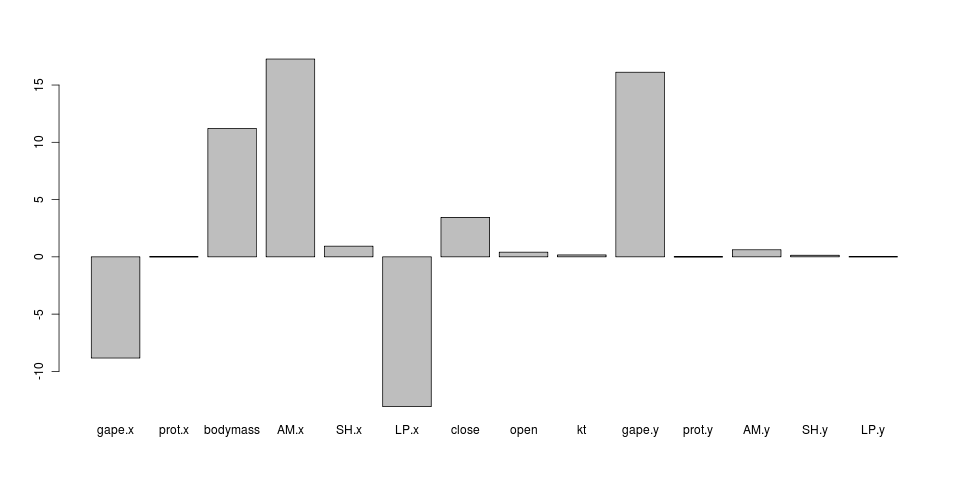
hmm, doesn’t match the general model fits: 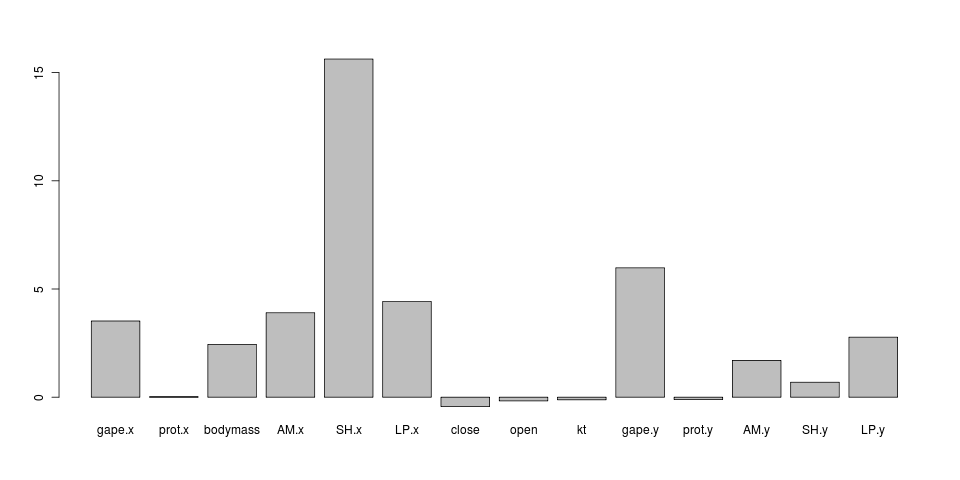
Puzzles and puzzle-shooting
A couple traits don’t converge for the general model:
conv(test) [1] “trait AM.x regime intramandibular model wright didn’t converge” [1] “trait SH.y regime intramandibular model wright didn’t converge” [1] “trait LP.y regime intramandibular model wright didn’t converge”
Above uses equal-alpha always as starting conditions (taken from hansen fit). Comparing against a fit attempt using very different alphas for each regime (intra at 0.1, other at 5). Comparisons run and pushed to flickr, gallery below. Results are clearly different; i.e. alpha ratios on release (alpha varies) model:
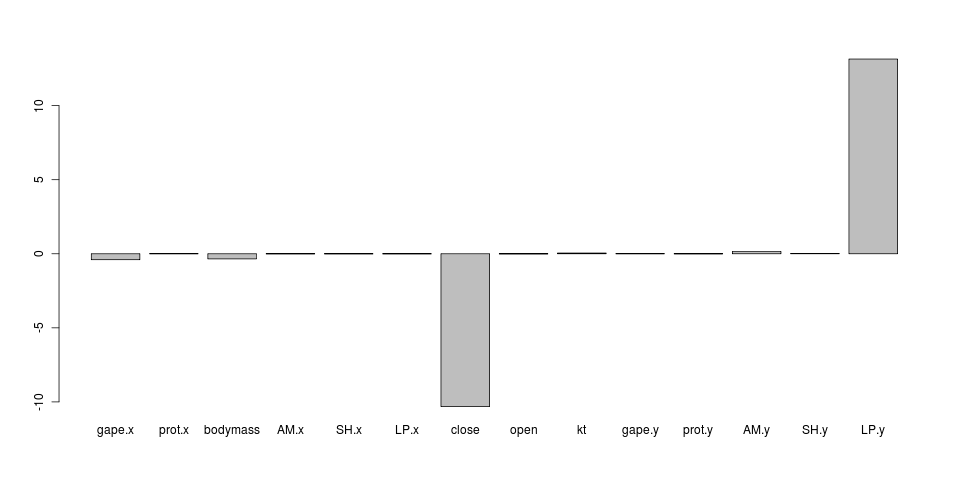
(replotted without the two dominant ratios here, first in stream below.)
[flickr-gallery mode=“search” tags=“phylogenetics” min_upload_date=“2011-05-05 18:00:00” max_upload_date=“2011-05-05 18:45:00”]
Results in more convergence errors, but not enough to explain difference (only one release_constraint model didn’t converge):
conv(test) [1] “trait bodymass, regime intramandibular, model ouch didn’t converge” [1] “trait bodymass, regime intramandibular, model release_constraint didn’t converge” [1] “trait LP.x, regime intramandibular, model ouch didn’t converge” [1] “trait open, regime intramandibular, model ouch didn’t converge” [1] “trait kt, regime intramandibular, model ouch didn’t converge” [1] “trait LP.y, regime intramandibular, model wright didn’t converge”
(commitwith this version of code)
Illustrates need for more robust searching of the likelihood surface!
Trying now with simulated annealing, shows a release of constraint:
wright_alphas intramandibular other gape.y 3.786699 12.59779 prot.y 9.275002 15.27857 release_alphas intramandibular other gape.y 2.008825 2.262495 prot.y 23.093936 25.768637
Protrusion strongly favors alpha model:
lliks $gape.y gen alpha sigma 0.0000000 0.7693642 1.0315282
$prot.y sigma gen alpha 0.00000 13.04190 13.44653
Worrisome that gen doesn’t do best though, it probably needs a slower cooling schedule for so many parameters. (code)
Trying with slower cooling, starting with unfavorable init conditions (higher alpha for intramandibulars):
release_alphas intramandibular other gape.y 1.969489 0.6064478 prot.y 25.087304 27.2187761 > wright_alphas intramandibular other gape.y 2.198133 4.446376 prot.y 20.443740 22.325272 >
Clearly just need super aggressive parameters. Robust search for actual inferences coming up. Need to add automated clever convergence testing.
Meetings:
Duncan
Discussing web tools to interface with APIs, with implementations in R. OAuth, CURL, XML, JSON. Approaches in REST vs SOAP. Notes (in prep, will be separate post.)